Browndye
Further Information
Other applications using Browndye
- Simulation-Enabled Estimation of Kinetic Rates (SEEKR).
Gallery
Here are some images from scientific projects using Browndye.
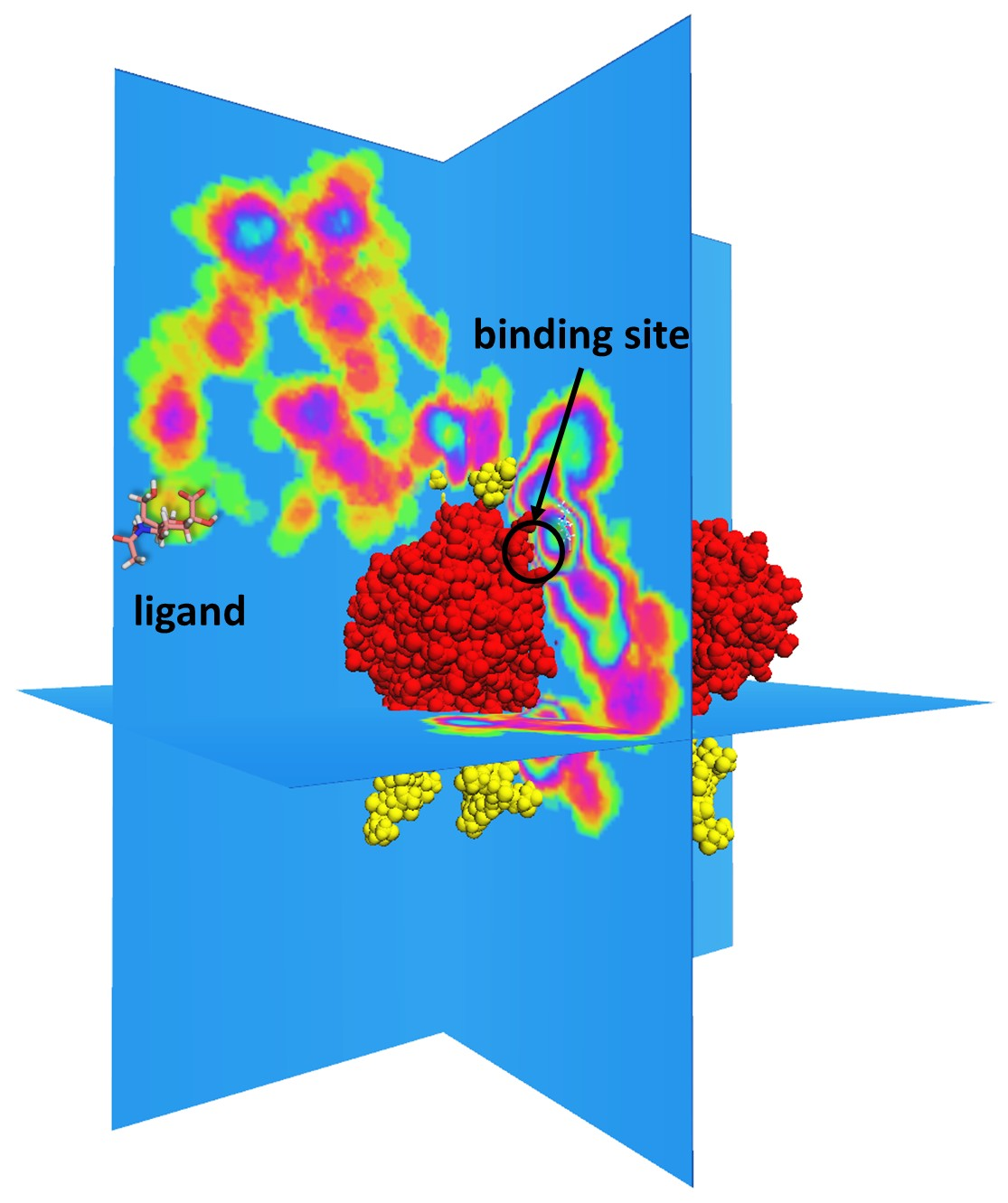
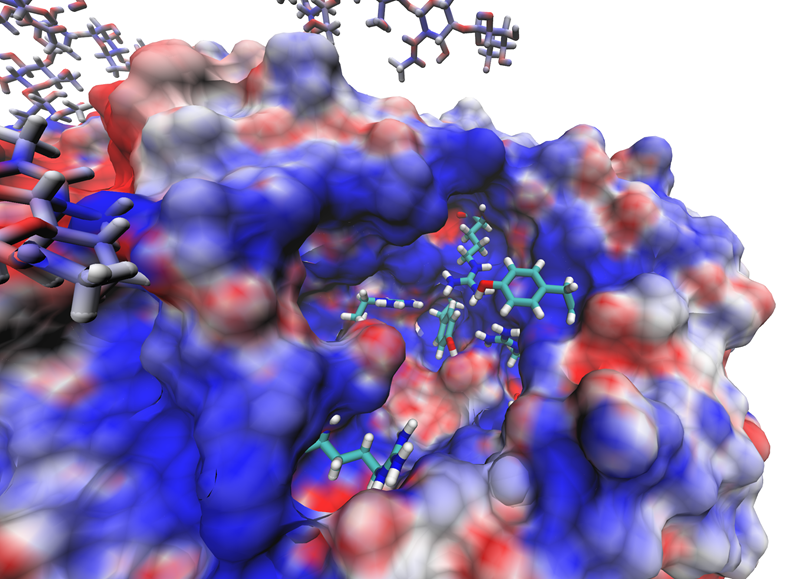
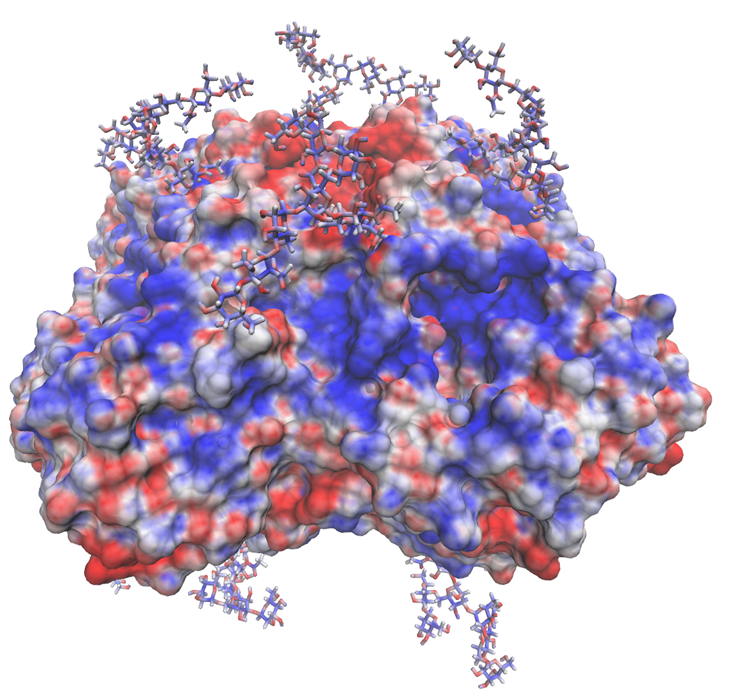
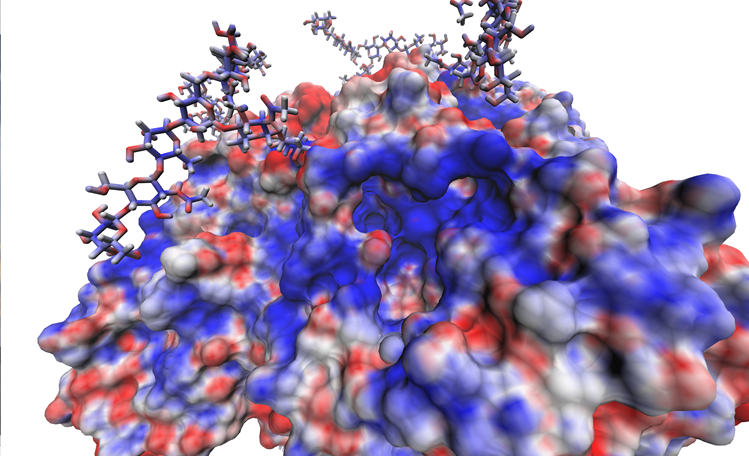
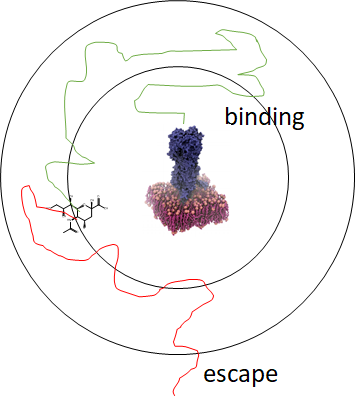
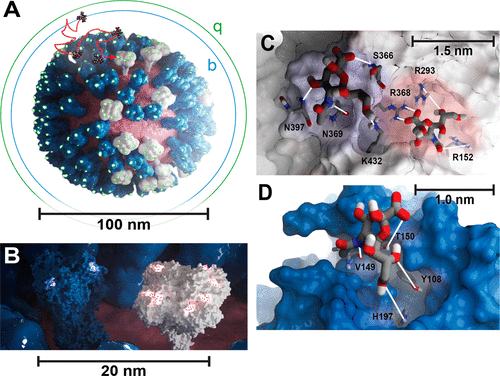
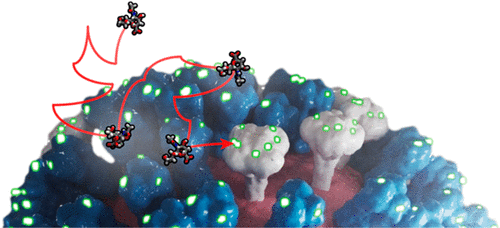
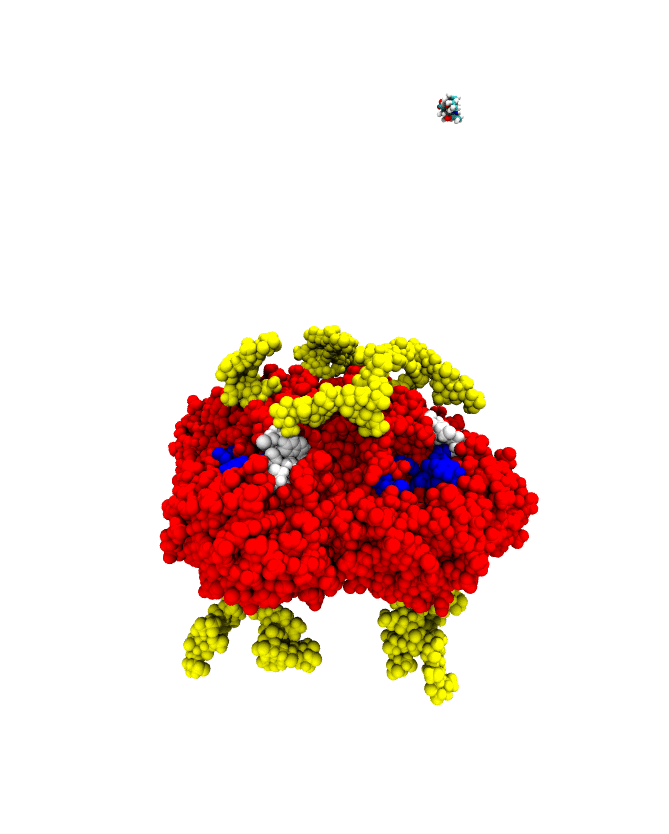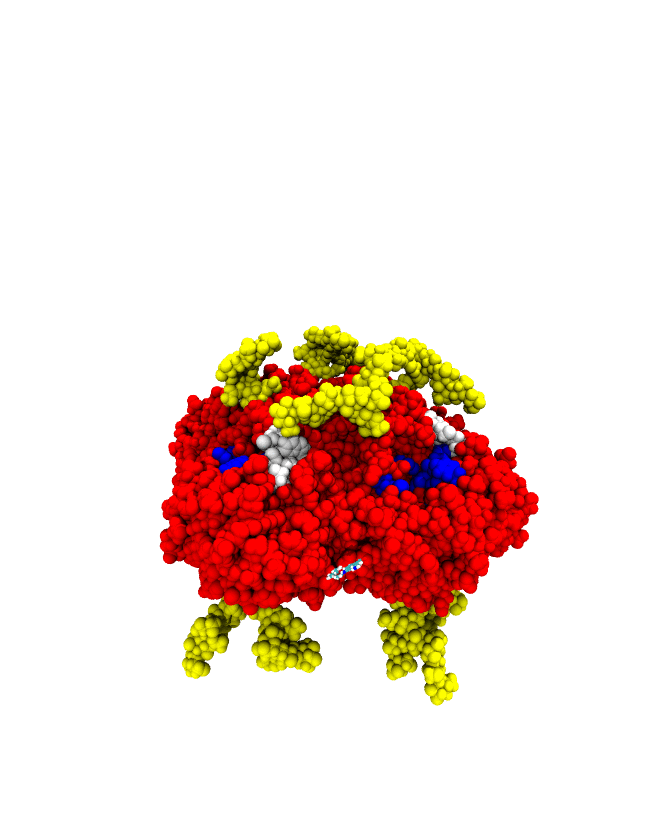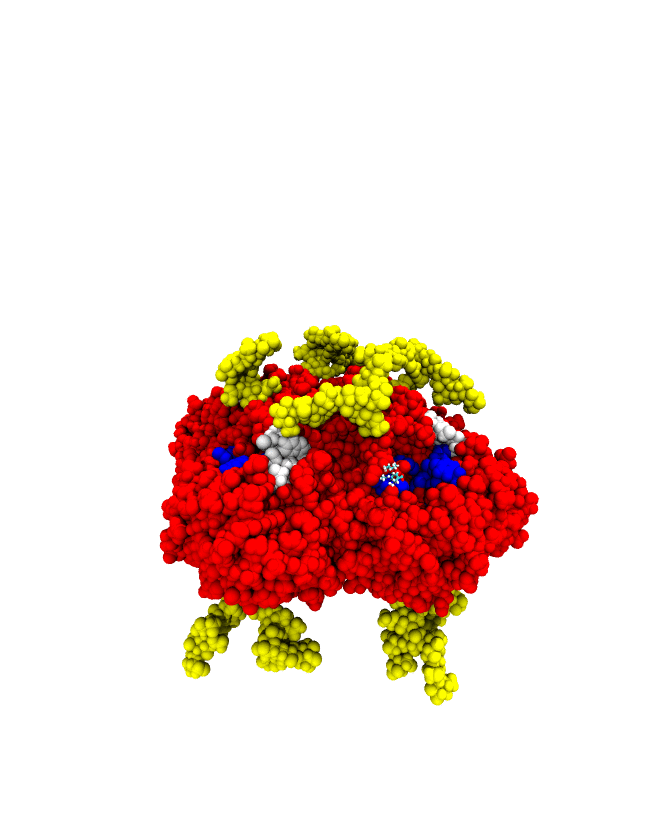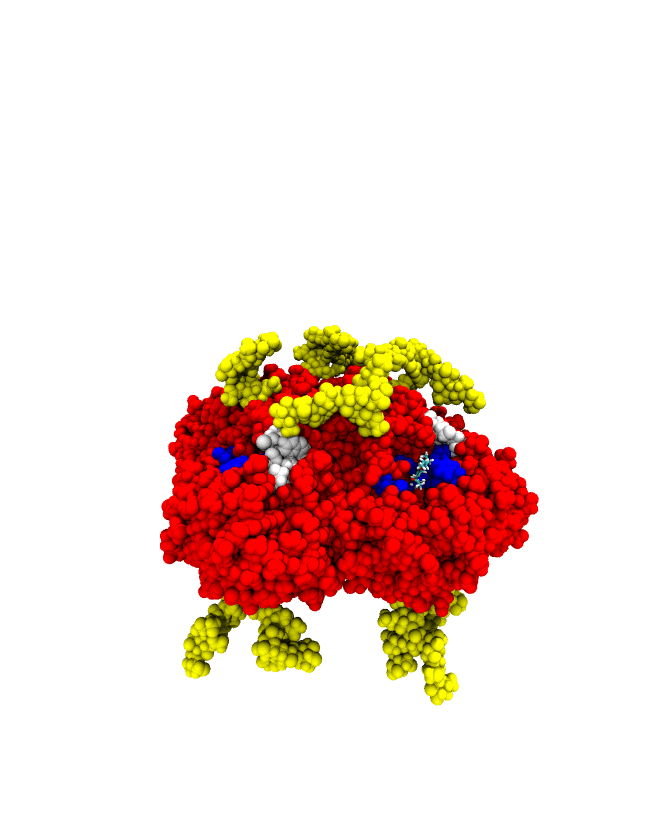
A sample successful trajectory, showing that the ligand explores significant space before finding the binding site. This sample NA contains glycans from the Glyprot server. Sialic acid (not to scale) is superimposed on the left side of the image for clarity. Seitz et. al. Biophysical Journal, 119, 2275-2289, 2020.
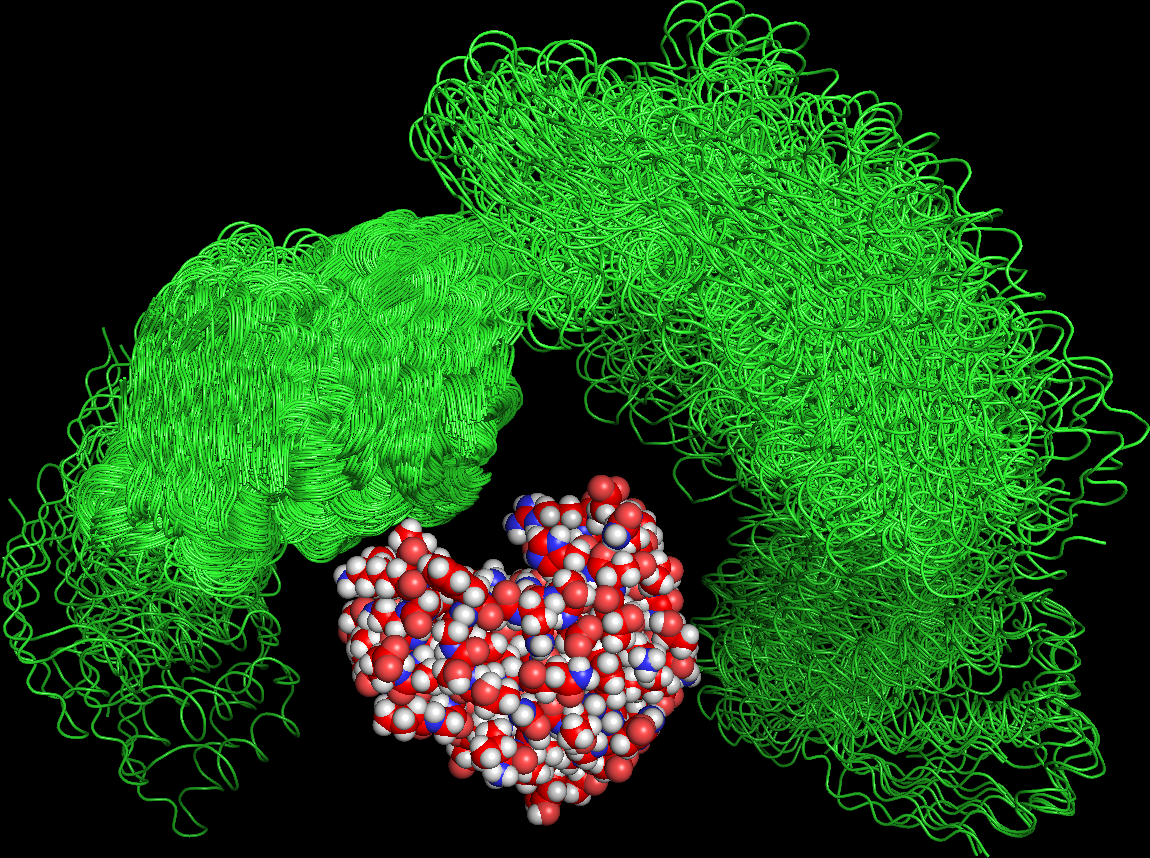
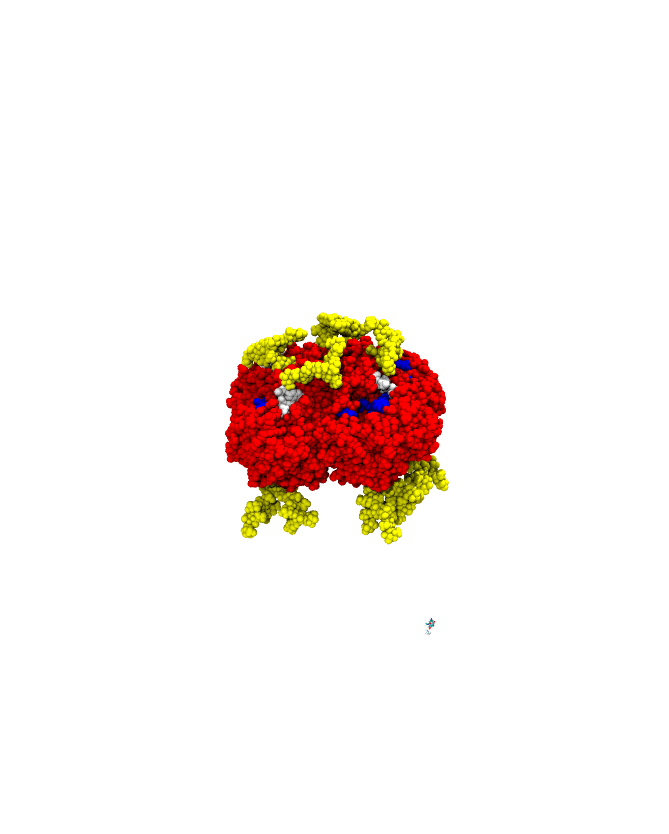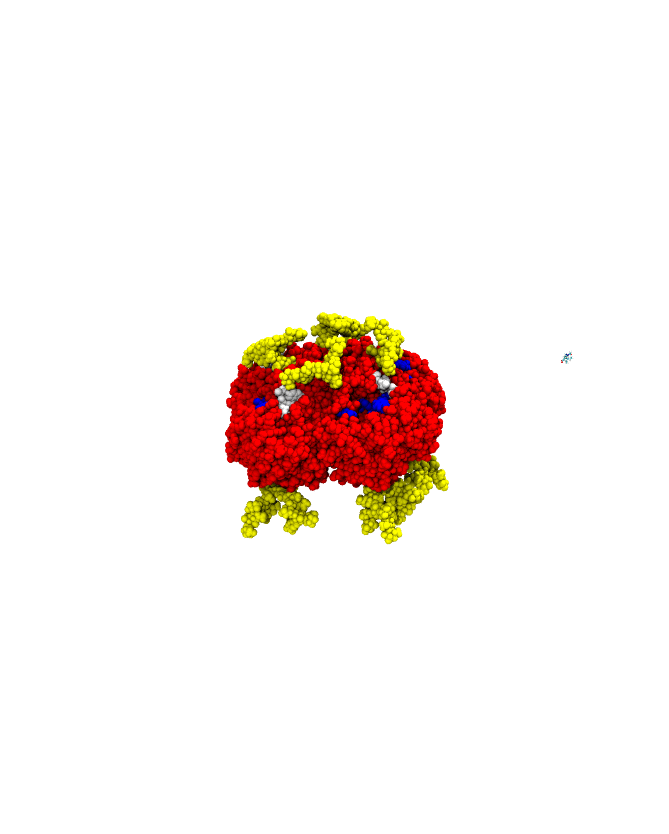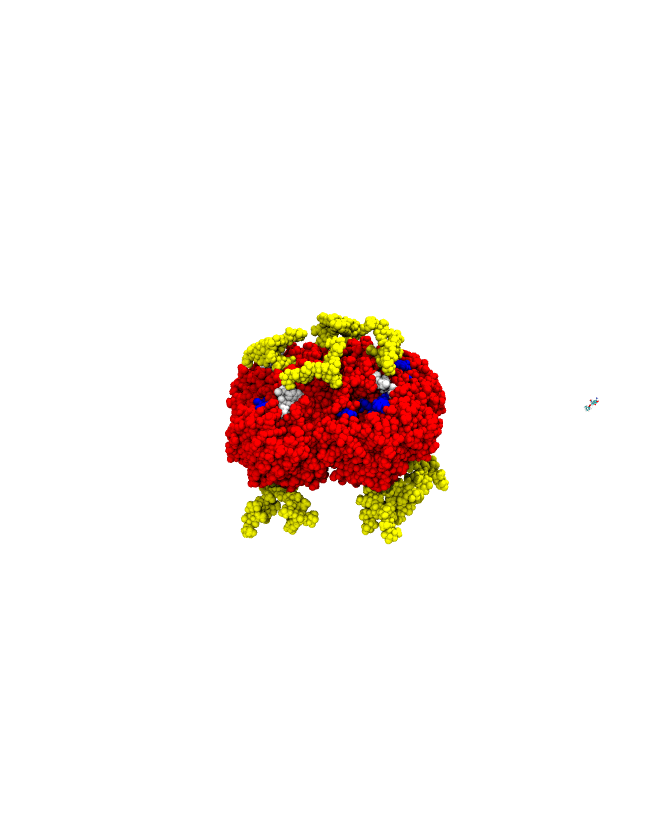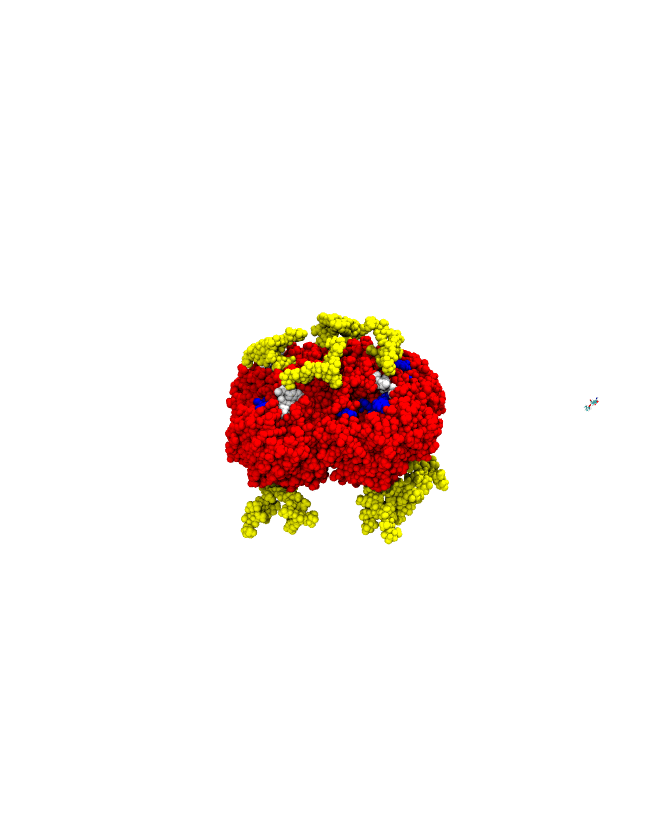
Barnase-barstar complex (1BRS) sample trajectory (R. Konecny/Pymol).
Votapka, Lane W., and Rommie E. Amaro. "Multiscale estimation of binding kinetics using brownian dynamics, molecular dynamics and milestoning." PLoS Comput Biol 11.10 (2015): e1004381.
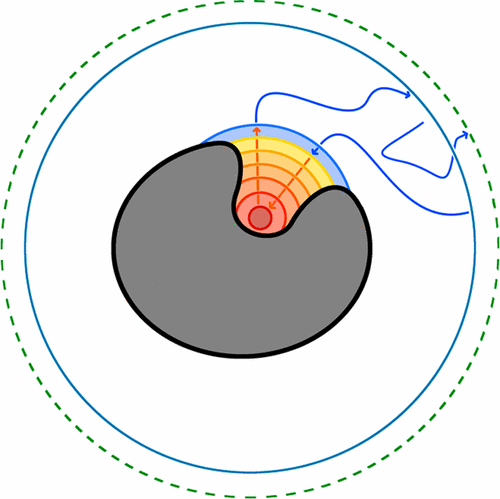
Votapka, Lane W., et al. "SEEKR: simulation enabled estimation of kinetic rates, a computational tool to estimate molecular kinetics and its application to trypsin–benzamidine binding." The Journal of Physical Chemistry B 121.15 (2017): 3597-3606.